Touching proteins with virtual bare hands: how to visualize protein-drug complexes and their dynamics in virtual reality
PubDate: Oct 2017
Teams: University of Warwick
Writers: Erick Martins Ratamero, Dom Bellini, Christopher G. Dowson, Rudolf A. Roemer
Abstract
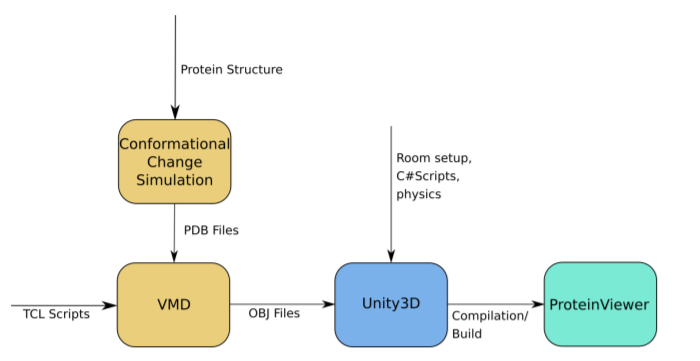
The ability to precisely visualize the atomic geometry of the interactions between a drug and its protein target in structural models is critical in predicting the correct modifications in previously identified inhibitors to create more effective next generation drugs. It is currently common practice among medicinal chemists while attempting the above to access the information contained in three-dimensional structures by using two-dimensional projections, which can preclude disclosure of useful features. A more precise visualization of the three-dimensional configuration of the atomic geometry in the models can be achieved through the implementation of immersive virtual reality (VR). In this work, we present a freely available software pipeline for visualising protein structures through VR. New customer hardware, such as the HTC Vive and the Oculus Rift utilized in this study, are available at reasonable prices. Moreover, we have combined VR visualization with fast algorithms for simulating intramolecular motions of protein flexibility, in an effort to further improve structure-lead drug design by exposing molecular interactions that might be hidden in the less informative static models.